How to use the Essential Genes database
The Essential Genes database integrates mice viability and phenotyping data and human essentiality data for a set of human and/or mouse genes, at users’ request. Below you can find information about the human and mouse gene attributes we are providing.
For context on this project, you may want to visit the IMPC Essential Genes landing page and our news post on essential genes.
To access the data you need to use our Hasura console. Ideally you need to have GraphQL knowledge but we have created 2 starter queries to get the basic information.
Step-by-step guide
1. Provide human genes or mouse genes
- Please use HGNC or MGI identifiers
- Format your list like this: “MGI:1929293”, “MGI:1098687” or “HGNC:19679”, “HGNC:24579”
Helper tools are available to find approved HGNC IDs or MGI IDs:
- You can use gene symbols as input and search for matches against approved gene symbols, synonyms, previous and withdrawn symbols.
- The following helper tools provide most recently approved gene symbols as well as HGNC or MGI identifiers:
- For human genes, the multi-symbol checker developed by HGNC: https://www.genenames.org/tools/multi-symbol-checker/
- For mouse genes, the batch query functionality developed by MGI: http://www.informatics.jax.org/batch
Data source:
- For human genes: https://www.genenames.org/download/statistics-and-files/
- For mouse genes: http://www.informatics.jax.org/downloads/reports/index.html
2. Choose your query
We have 2 starter queries, one to get human gene orthologs and details by providing a list of MGI IDs:
query getDetailsByMGIAccessionID {
mouse_gene(where: {mgi_gene_acc_id: {_in: [LIST_OF_IDS]}}) {
mgi_gene_acc_id
symbol
ensembl_gene_acc_id
mgi_chromosome
mgi_start
mgi_stop
subtype
type
orthologs {
human_gene {
symbol
hgnc_acc_id
human_gene_synonym_relations {
human_gene_synonym {
synonym
}
}
hgnc_genes {
location_sortable
locus_group
}
achilles_gene_effects {
mean_gene_effect
entrez_acc_id
id
}
}
is_max_human_to_mouse
is_max_mouse_to_human
support
fusils {
bin
bin_code
id
}
}
mouse_gene_synonym_relations {
mouse_gene_synonym {
synonym
}
}
}
}And another to get mouse gene orthologs and details with a list of HGNC IDs:
query getDetailsByHumanGeneIDs {
human_gene(where: {hgnc_acc_id: {_in: [LIST_OF_IDS]}}) {
hgnc_acc_id
ensembl_gene_acc_id
name
symbol
orthologs {
mouse_gene {
symbol
type
subtype
mouse_gene_synonym_relations {
mouse_gene_synonym {
synonym
}
}
}
support
is_max_human_to_mouse
is_max_mouse_to_human
fusils {
bin
bin_code
id
}
}
hgnc_genes {
location_sortable
locus_group
}
achilles_gene_effects {
mean_gene_effect
entrez_acc_id
human_gene_id
id
}
}
}Copy the query you are interested, replace LIST_OF_IDS with your list and copy it to the Hasura console, it will look like this:
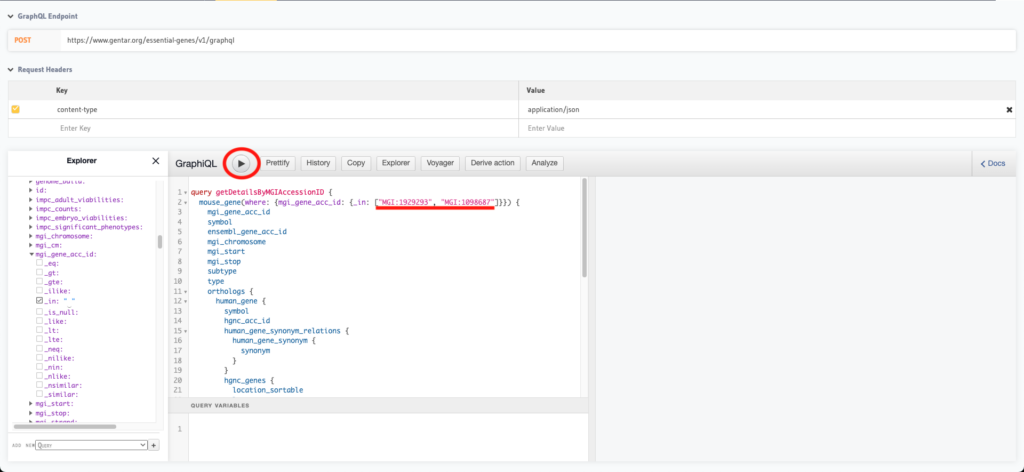
3. Execute the query and get the data
The Hasura console is straightforward to use, after the query has been copied, you need only to click in the “Execute Query” button and you will get all the available data. The data is also available from this GraphQL endpoint: https://www.gentar.org/essential-genes/v1/graphql
Data available
1. Human gene attributes:
1.1. Human cell essentiality scores – Cancer Dependency Map, Achilles gene effect.
For each gene, we compute and provide the mean gene effect across all cell lines in the corresponding data release (for example, for data release 20Q1, the data involved 739 cell lines for 18,333 genes).
- Gene identifiers: Entrez IDs, as provided in Achilles file.
- Lower values indicate more intolerance (i.e. more essential).
- We used the threshold -0.45 to classify genes in Essential (below the threshold) and Non-Essential (above the threshold), as per our IMPC paper in Nat Comm 2020.
- [ Data source: https://depmap.org/portal/download/ ]
1.2. Constraint scores – gnomAD v2, pLoF Metrics by Gene TSV.
- Gene identifiers: Ensembl IDs or gene symbols, as provided in the gnomAD file.
- [ Data source: https://gnomad.broadinstitute.org/downloads#v2-constraint ]
1.3. Gene Dosage sensitivity –The ClinGen Dosage Sensitivity curation process collects evidence supporting/refuting the haploinsufficiency and triplosensitivity of genes and genomic regions.
- Gene identifiers: symbols, as provided by ClinGen.
- [ Data source: https://search.clinicalgenome.org/kb/gene-dosage ]
1.4. Illuminating the Druggable Genome – IDG classification of genes in Tdark, Tbio, Tchem, Tclin.
- [ Data source: https://druggablegenome.net ]
2. IMPC gene attributes:
2.1. IMPC primary (adult) viability – The IMPC conducts an early adult (pre-weaning) viability screen to determine gene essentiality (analysed as defined in IMPRESS). Briefly:
- A minimum of 28 pups are genotyped.
- The line is classified as:
- Lethal: absence of live knockout (null) homozygote pups.
- Subviable: fewer than 12.5% live homozygous pups (half of the 25% expected; P < 0.05, binomial distribution).
- Viable: homozygous (null and wild type) and heterozygous pups are observed in the same or more than the expected normal Mendelian ratios.
- A viable call is also made when there are less than 28 total pups and homozygous null pups ≥ 4 (as this would result in ≥ 14% homozygous pups when 28 pups were genotyped).
- The distribution of lines into these 3 categories remains almost the same across data releases (lethal 24%, subviable 9% and viable 67% for DR 9.1).
- Gene identifiers: MGI IDs.
- [ Data source: https://www.mousephenotype.org ]
3. FUSIL bin categorization – IMPC viability and phenotype data and mean Achilles gene effect scores are combined to define one of five mutually exclusive categories, as per our IMPC paper on essential genes, Nat Comms 2020. The following categories are defined:
- Cellular Lethal (CL): IMPC lethal and mean Achilles gene effect ≤ -0.45;
- Developmental Lethal (DL): IMPC lethal and mean Achilles gene effect > -0.45;
- Subviable (SV): IMPC subviable and mean Achilles gene effect > -0.45;
- Viable with Phenotype (VP): IMPC viable and mean Achilles gene effect > -0.45, and with at least one IMPC significant phenotype;
- Viable No Phenotype (VN): as above, but no IMPC significant phenotype (when at least 13 or more phenotype procedures have been analysed for the homozygous (null) allele knockouts of the corresponding mouse line).
Other categories (only a few genes are found to be in any of these situations):
- V.insuffProcedures: as above for viable, but less than 13 phenotype procedures analysed so far
- SV.outlier: subviable and with mean Achilles gene effect ≤ -0.45
- V.outlier: viable and with mean Achilles gene effect ≤ -0.45
We integrate using orthologues
Orthologue inferences are based on the HCOP assertions, based on 12 algorithms (see here for more on the HCOP tool, developed by the HGNC). We determine the confidence on the orthologue prediction based on the number of methods that support each inference:
- 12-9 methods (100-75%) support the same inference: good-confidence orthologue;
- 8-5 methods (67-42%): moderate-confidence orthologue;
- 4-1 methods (33-8%): low-confidence orthologue;
- 0 methods: no orthologue.
We provide FUSIL bins when:
- the orthologue inference is supported by ≥ 5 methods;
- the score is maximum in both directions, mouse-to-human and human-to-mouse (i.e. we filter out genes with duplicated maximum scores or no maximum scores in both directions).
