The IMPC consortium uses different targeting strategies that have different and complementary properties to produce knockout alleles. These strategies rely on the identification of a ‘critical’ exon common to all transcript variants that, when deleted, disrupts gene function. The mutant alleles may include additional functional features, such as a lacZ reporter cassette or conditional potential.
The IMPC has been increasingly using CRISPR/Cas9 to knockout genes for several years now. Prior to this, gene knockout technology was based on ES cells. Here Lauryl Nutter talks about CRISPR/Cas9, its uses for studying mouse genetics and how the technique is being used by the IMPC.
Below is a general outline of all the targeting strategies that are used or have been used by IMPC teams.
CRISPR/Cas9 design
IMPC alleles viable for high throughput pipeline
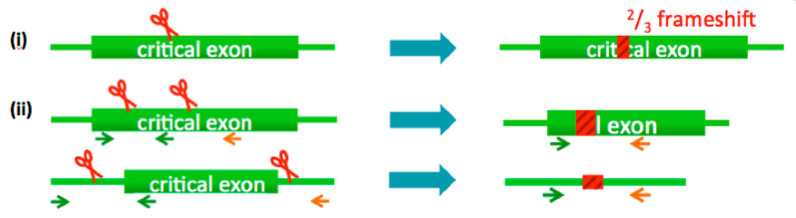
(i) Small deletions (single cut strategy)
(ii) Exon deletion / large deletions (2 cut strategy)
Alleles on request
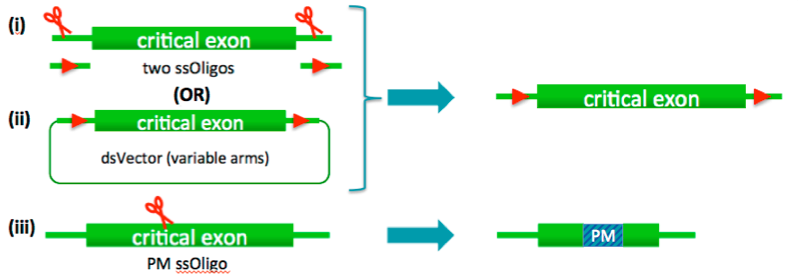
(i) LoxP-flanked critical regions
(ii) Point mutations
(iii) Conditional and lacZ reporter allele – insertion of dsVector
‘Knockout-first’ design
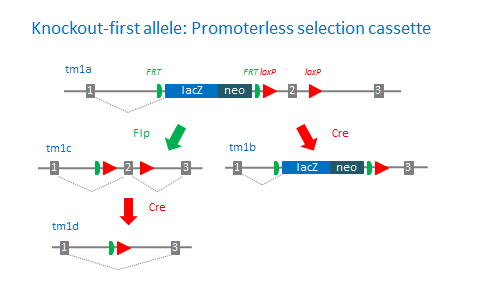
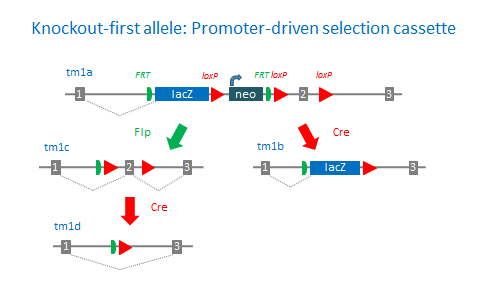
The ‘knockout-first’ allele design is a flexible strategy that can be used to produce reporter knockouts, conditional knockouts, and null alleles following exposure to site-specific recombinases Cre and Flp.
Promoterless and promoter-driven targeting cassettes are used for the generation of a ‘knockout-first’ allele (tm1a) in C57BL/6N embryonic stem cells (Pettitt et al., 2009). These two strategies have been shown to yield different targeting efficiencies (Skarnes et al., 2011). Insertion of a lacZ trapping cassette and a floxed promoter-driven neo cassette inserted into the intron of a gene is expected to disrupt gene function. Flp converts the ‘knockout-first’ allele to a conditional allele (tm1c), restoring gene activity. Cre deletes the promoter-driven selection cassette and floxed exon of the tm1a allele to generate a lacZ– tagged allele (tm1b) or deletes the floxed exon of the tm1c allele to generate a frameshift mutation (tm1d).
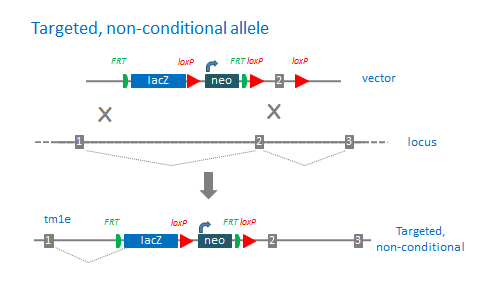
‘Targeted, non-conditional’ alleles (tm1e) are missing the downstream loxP site. The 3’ loxP site is often lost due to recombination events in the homology region between the targeting cassette and 3’ loxP site. These mutations cannot be converted to conditional alleles following Flp treatment.
Standard deletion alleles with the promoter-driven targeting cassette:
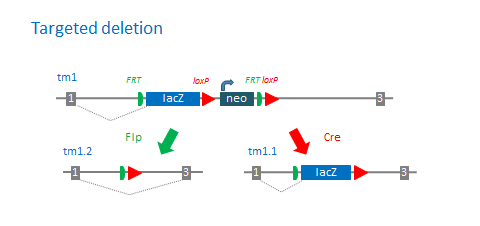
Number of available derivative alleles:
- tm1a: KO first allele (reporter-tagged insertion allele): 0
- tm1b: Reporter-tagged deletion allele (post-Cre): 0
- tm1c: Conditional allele (post-Flp): 0
- tm1d: Deletion allele (post-Flp and Cre with no reporter): 0
- tm1e: targeted, non-conditional allele: 0
- tm1: Reporter-tagged deletion allele (with selection cassette): 0
- tm1.1: Reporter-tagged deletion allele (post Cre, with no selection cassette): 0
- tm1.2: Reporter-tagged deletion allele (post Flp, with no reporter and selection cassette): 0
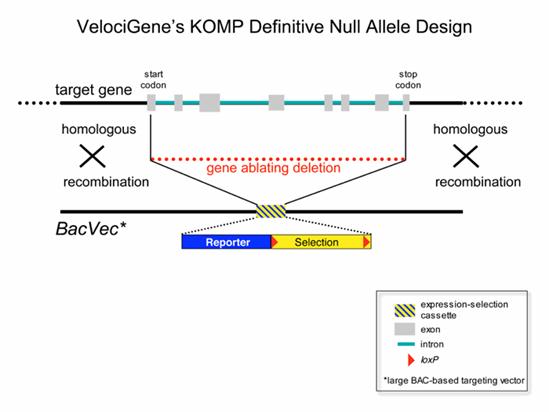
Velocigene null allele design
In most cases this design will result in complete null alleles that delete the entire protein coding sequence of the target gene. This allele design can be applied to any gene transcribed by RNA polymerase II regardless of its size, intron-exon structure, RNA splicing pattern, or protein-coding capacity (Valenzuela et al., 2003).